Drugging the Undruggable: How PROTACs Hijack Cellular Machinery to Destroy Cancer
Drugging the Undruggable: How PROTACs Hijack Cellular Machinery to Destroy Cancer
Daniil Prokofev
Thomas Jefferson High School for Science and Technology
This article placed 3rd in the 2025 Teknos Writing Contest.
Cancer is the second leading cause of death in the U.S., with drug resistance and the resulting ineffectiveness of following drug treatment being responsible for 90% of cancer related deaths [7]. Most anticancer drugs are small molecules that inhibit cancer-driving proteins. As a result, cancer cells, driven by relentless evolutionary pressure from those drugs, simply overproduce the target protein, manufacturing so many that the drug is no longer effective. To that point, an estimated 85% of disease-causing proteins in the human body are considered “undruggable” since they lack well-defined active sites for inhibitors to bind to [5]. This gives oncology two major questions: How can we prevent or revert drug resistance where cells just overproduce targets? And how can we attack the vast majority of cancer-causing proteins that we seemingly can’t target?
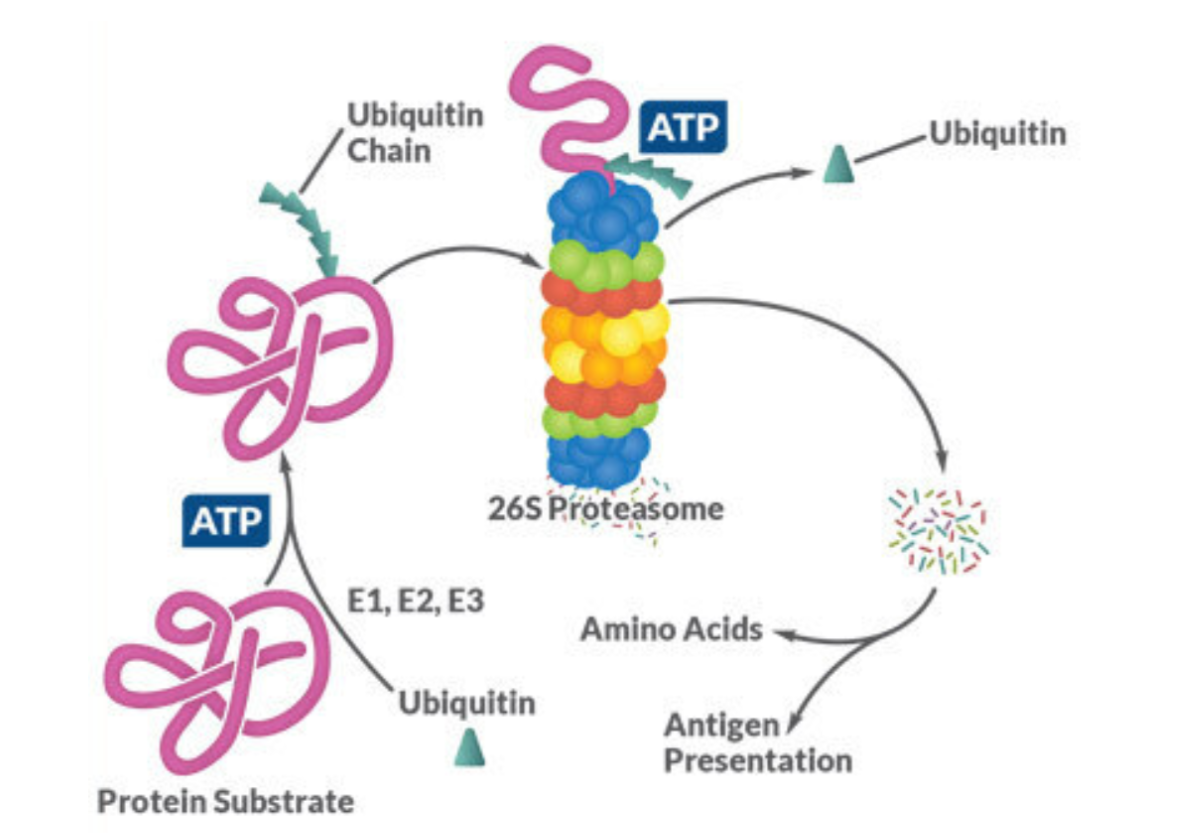
This is where PROTACs come in. Proteolysis Targeting Chimeras (PROTACs) are small, innovative molecules that do not inhibit their targets, but rather hijack the cell’s own garbage disposal—the Ubiquitin-Proteasome System (UPS). Every cell uses the UPS to dispose of old, unneeded or damaged proteins. This process works via a “tag and destroy” mechanism. An enzyme called E3 ligase identifies a target protein and attaches a molecular “tag” called ubiquitin. This tag acts as a signal, marking the protein for degradation. A large protein complex, the proteasome, then recognizes the ubiquitin tag, unfolds the unneeded protein, and cuts it up into amino acids, recycling it for further use [2].
Figure 1. The Ubiquitin-Proteasome System (UPS). The UPS acts as the cell's natural garbage disposal, where E1, E2, and E3 enzymes work to tag a target protein with a ubiquitin chain. The 26S proteasome then recognizes this tag, degrades the protein into amino acids, and recycles it [2].
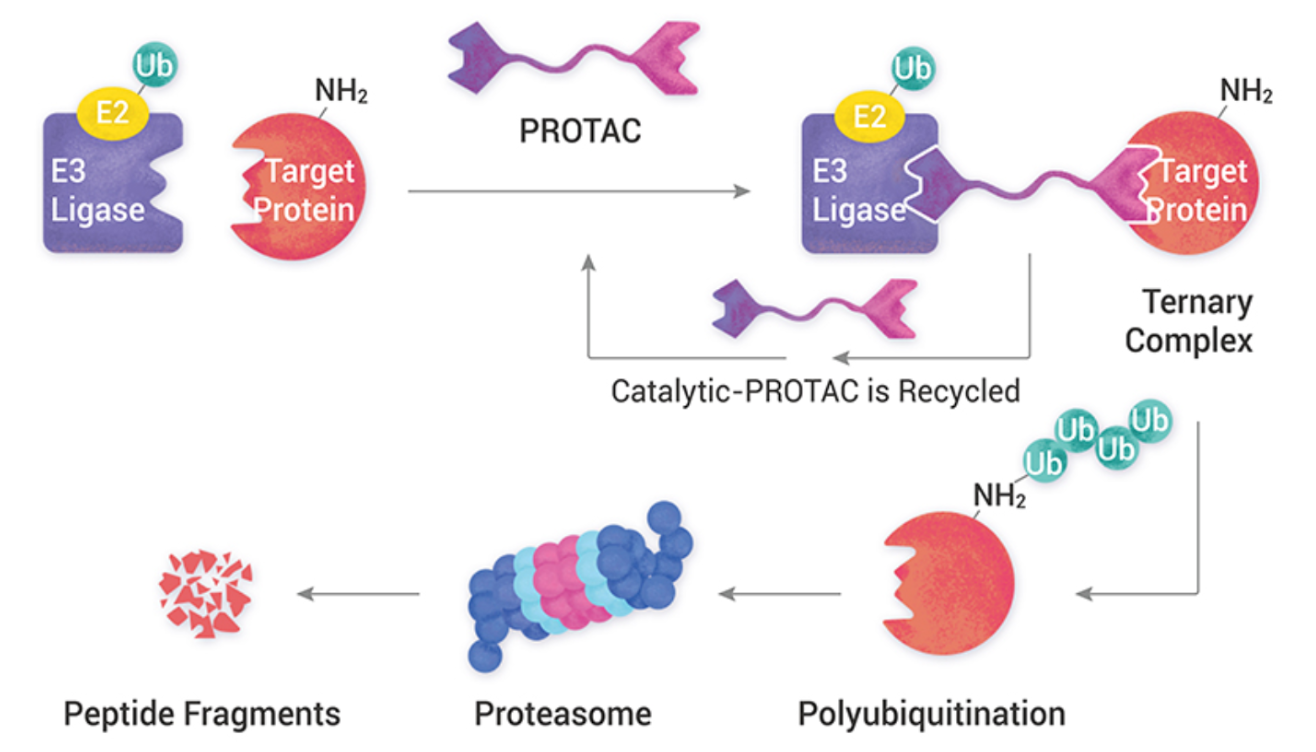
The PROTAC molecule elegantly takes over this process. PROTACs are “chimeric” molecules with two distinct “heads” connected by a flexible linker. One head is engineered to bind to a target cancer protein, the Protein of Interest (POI). The other head is designed to grab onto an E3 ligase. By simultaneously binding to both, the PROTAC physically brings the cancer protein close to the UPS, forcing the E3 ligase to tag the protein with ubiquitin, with the proteasome doing the disposal and destroying the protein. After the protein’s destruction, the PROTAC molecule is released and proceeds to find another target to dispose of. PROTACS have a major advantage over traditional inhibitors in this sense, since they are not a one-time-use molecule [6].
Figure 2. The PROTAC mechanism. A PROTAC molecule binds to both a target protein and an E3 ligase, forming a ternary complex. This proximity allows the E3 ligase to tag the protein with ubiquitin (polyubiquitination), marking it for destruction. The proteasome then degrades the protein into peptide fragments, and the catalytic PROTAC is recycled [6].
The exciting potential of PROTACs is quite obvious, but the scientific literature gives light to both successes and significant challenges of this new biotechnology. On one hand, PROTACs can overcome drug resistance and cancer proliferation driven by protein overexpression, without depletion of the agent. This has been demonstrated in clinical trials for prostate cancer, where the PROTAC drug ARV-110 successfully degrades the AR protein in patients who had progressed on conventional inhibitors, leading to tumor shrinkage [3]. Equally important, since PROTACs only need a small binding pocket anywhere on the protein, they demonstrate successful degradation of “undruggable” targets in preclinical studies [1].
PROTACs are not a silver bullet and come with their own set of challenges, however. They demonstrate a counterintuitive phenomenon known as the “hook effect,” where at very high concentrations, the drug becomes less effective. This occurs because the excess PROTAC molecules bind to target proteins and E3 ligases separately, preventing the assembly of the three part “ternary complex” needed for degradation [4]. Furthermore, the actual engineering of these large PROTAC molecules is quite challenging, since formulating them into a stable pill with good oral bioavailability is difficult [6]. Recent studies have also shown that cancer cells can develop resistance to PROTACs by mutating the binding site on the target protein or by mutating the E3 ligase the PROTAC relies on, effectively disabling its mechanism [8].
Despite these challenges, targeted protein degradation is revolutionary. No matter how you put it, PROTACs are making undruggable cancer proteins druggable and are working in clinical studies where no other drug could [3]. The core concept has already inspired the next generation of therapeutic platforms, such as LYTACs, which use a similar principle to destroy disease-causing proteins outside of the cell [6]. By hijacking the cell’s own natural and essential mechanisms to destroy cancer-critical proteins, these molecular therapies will aid significantly in the fight against cancer.
References
[1] Békés, M., Langley, D. R., & Crews, C. M. (2022). PROTAC targeted protein degraders: the past is prologue. Nature Reviews Drug Discovery, 21(3), 181–200. https://doi.org/10.1038/s41573-021-00371-6
[2] Földesi, B. (2019, February 3). Recycling the Cell: Autophagy and the Ubiquitin-Proteasome Processes. Biomol Blog. https://resources.biomol.com/biomol-blog/recycling-the-cell-autophagy-and-the-ubiquitin-proteasome-processes
[3] Gao, X., Burris III, H. A., Vuky, J., Dreicer, R., Sartor, A. O., Sternberg, C. N., ... & Petrylak, D. P. (2022). Phase 1/2 study of ARV-110, an androgen receptor (AR) PROTAC degrader, in metastatic castration-resistant prostate cancer (mCRPC). Journal of Clinical Oncology, 40(6_suppl), 17. https://doi.org/10.1200/JCO.2022.40.6_suppl.017
[4] Li, K., & Crews, C. M. (2022). PROTACs: Past, Present and Future. Chemical Society Reviews, 51(12), 5214-5236. https://pmc.ncbi.nlm.nih.gov/articles/PMC10237031/
[5] Lu, Y., Yang, Y., Zhu, G., Zeng, H., Fan, Y., Guo, F., ... & Ge, G. (2023). Emerging Pharmacotherapeutic Strategies to Overcome Undruggable Proteins in Cancer. International Journal of Biological Sciences, 19(11), 3360–3382. https://pmc.ncbi.nlm.nih.gov/articles/PMC10367563/
[6] MedChemExpress. (n.d.). PROTAC — Future of Drug Molecules from Modular Construction. https://www.medchemexpress.com/literature/protac-future-of-drug-molecules-from-modular-construction.html
[7] Wang, X., Zhang, H., & Chen, X. (2019). Drug resistance and combating drug resistance in cancer. Cancer Drug Resistance, 2(2), 141-160. https://doi.org/10.20517/cdr.2019.10
[8] Zhang, L., Riley-Gillis, B., Vijay, P., & Shen, Y. (2019). Acquired Resistance to BET-PROTACS (Proteolysis-Targeting Chimeras) Caused by Genomic Alterations in Core Components of E3 Ligase Complexes. Molecular Cancer Therapeutics, 18(7), 1302–1311. https://doi.org/10.1158/1535-7163.mct-18-1129
[9] Zhong, L., Li, Y., Xiong, L., Wang, W., Wu, M., Yuan, T., ... & Yang, S. (2021). Small molecules in targeted cancer therapy: advances, challenges, and future perspectives. Signal Transduction and Targeted Therapy, 6(1), 201. https://doi.org/10.1038/s41392-021-00572-w