An Unexpected Role In Cell Adhesion
Spindle-Associated Membrane Protein 1 (SAMP1) May Play an Unexpected Role In Cell Adhesion
Abhishek Mogili
Dr. Jeremy Smyth
Uniformed Services University of the Health Science
Thomas Jefferson High School for Science and Technology
Abstract
Previous cell biology studies have shown that the endoplasmic reticulum (ER) and microtubules (MTs) co-localize with each other during both interphase and mitosis. The protein SAMP1 may be the linking protein between the ER with MTs because it has been shown to localize with both ER membranes and tubulin. In this experiment RNAi treatments were used to stop SAMP1 synthesis in order to determine its function and analyze changes in cell morphology. Differences in cell structure were observed in HeLa cells notably with our second RNAi treatment. Cells began to present with stress fibers, and elongated cell structures, that were different from the normal HeLa cells we had observed previously. Mitotic cells also did not form perfect spheres, in that portions of the cell were often still attached to the coverslip. These findings indicate that the SAMP1 protein could play a major role in cell adhesion and structure.
Introduction
This past summer I had the opportunity to work with Dr. Jeremy Smyth and his lab at the Uniformed Services University in Bethesda, MD. His lab is interested in learning more about the morphology and function of the ER which plays a vital role in Mammalian cells. Its wide range of functions include Lipid Synthesis, Protein regulation, and Calcium ion storage. Specifically my project looked at the relationship between the ER and microtubules, a structural protein that forms the cytoskeleton in most eukaryotic cells. Previous studies published by Dr. Smyth’s lab have shown that proteins that are found in the ER and proteins found in microtubules co-localize during all phases of cell growth and division indicating a possible relationship. Furthermore, a study published by Vijayaraghavan et al has shown that a relatively new protein Spindle-Associated Membrane Protein 1(SAMP1) also co-localizes with the ER and Microtubules. In order to determine its function and possible linkage, loss of function analysis was done by knocking out the gene that codes for it and determining the effects on cell structure that resulted. My project explores this relationship and attempts to form a linkage relationship between the two proteins. This research has implications in developmental biology and can be used to look at possible ER-related diseases like those related to metabolic diseases [2].
Methods
The primary method that this experiment used was Immunofluorescence microscopy. This includes a three-step procedure. The first step includes the treatment phase, where HeLa cells were cultured and subjected to an RNAi treatment to knockout the gene that codes for the SAMP1 protein. The cells were grown on glass coverslip slides in order to make it easy to prepare and mount for viewing. After the treatments, the second step involves fixing the HeLa cells with Glutaraldehyde, this protocol involved lysing the cells with Karsenti Buffer in order to “freeze” the organelles. The last step includes the immunofluorescence staining using a two-step process. We used antibodies that bind to proteins specific to the ER and Microtubules. For the ER we used an antibody that binds to Calreticulin and one that binds to PDI both found in the ER, and for Microtubules we used an antibody that binds to Tubulin. We also used DAPI to stain for DNA and is shown in blue in the images. For our secondary antibodies we used antibodies created in Mice and Rabbits that bind to the primaries that we used. We also stained for DNA with DAPI. Due to the expensive nature of these antibodies, we conducted a few range-finding stains in order to determine the optimal concentration and company. This was done in order to ensure that there was enough antibody to stain all of the protein that is available and to prevent over-saturation. After staining all slides were looked at under a Nikon A1 confocal microscope. Images were taken and were the primary source of our data.
Data and Analysis
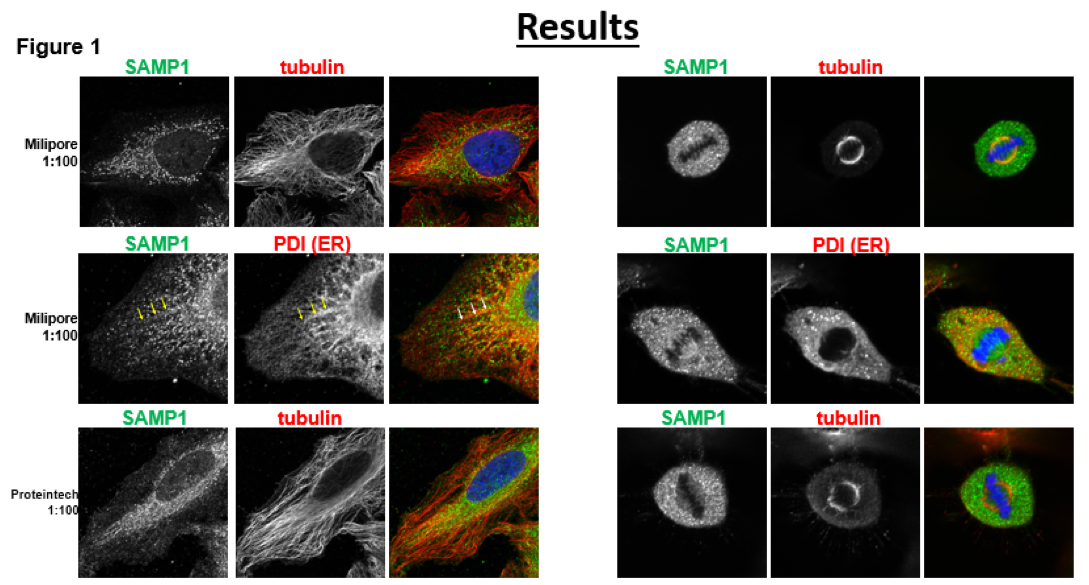
Part of this experiment involved the development of an effective procedure with the RNA Interference treatments. Our initial results aimed to replicate the results that were found by Buch et al. Our images initially didn’t show similar results, depicted to the right, the color image shows the overlay of the DNA(Blue), Endoplasmic Reticulum/SAMP1(green) and Microtubules(Red). From this initial study we can see that PDI (ER) and SAMP1 which is our target protein.
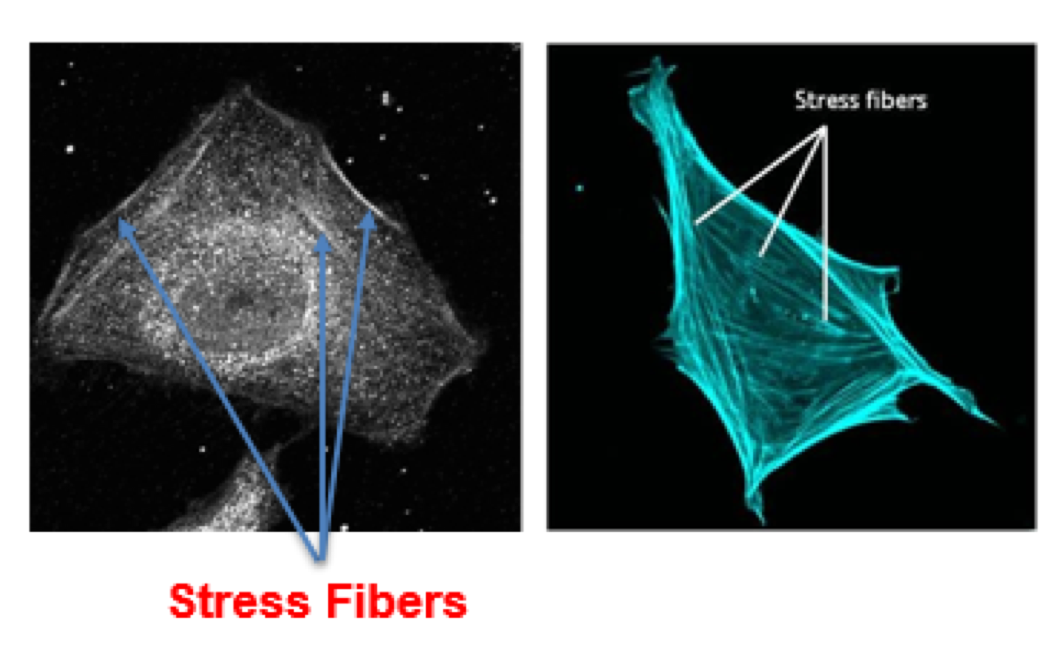
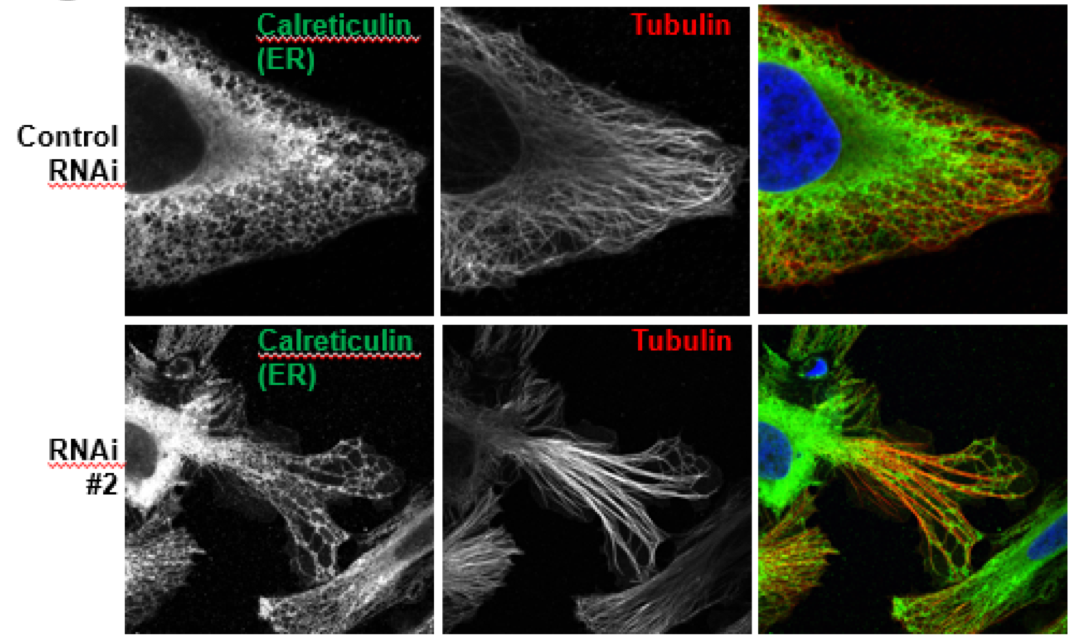
Although co-localization provides us some insight into the function of the protein, further tests must be done to establish a trend. Many protein function studies involve loss-of-function studies. We used an RNAi treatment that destroys the mRNA produced by the cell and therefore results in the non-expression of the SAMP1 protein. We then stained the cells using the same method and analyzed the changes in cell morphology. As shown in (Figure 2) the cells that were afflicted with the RNAi #2 treatment, showed drastic changes in the cytoskeletal structure. The long elongated structures are called stress fibers and consist of myosin and actin. These proteins and structures form when the cell undergoes extreme stress. This result was completely unexpected and showcases an interesting link between SAMP1 and the cell structure.
Conclusions
Although a clear definitive link was not established between SAMP1 and the co-localization of the Endoplasmic Reticulum and Microtubules, a promising beginning has been made in trying to determine the true effect SAMP1 has in the cell. The absence of SAMP1 resulted in elongated cytoskeletal structures that resemble actin-based stress fibers in both mitotic and interphase cells. This result draws us to believe that SAMP1 is an integral protein in cell adhesion and cytoplasmic structure. These unexpected results gave us a glimpse into the true function of the SAMP1 protein and its diverse role in the cell.
Discussion
The results that can be derived from this study revolve around the role that SAMP1 plays in the cell. We hypothesized that SAMP1 would form the link between Microtubules and the Endoplasmic Reticulum. Our results, however, indicated that SAMP1 may play an even more vital role in the cell and construct a vital role in cell adhesion and cytoskeleton formation.
Further studies will be conducted at the Uniformed Services University to confirm the RNAi treatment effectiveness, and to test different controls of gene expression, to truly obtain a loss of function analysis. Western blot protein analysis will be conducted to test the effectiveness of each RNAi treatment and to ensure that the protein is not expressed. Furthermore, tests will be conducted using different dilutions of antibodies and secondary staining methods in order to change the resolution of each image. This will hopefully result in an optimal saturation and provide us a reliable set of images from which to draw conclusions.
References
[1] Buch, C., Lindberg, R., Fiqueroa, R., Gudise, S., & Hallberg, E. (2009). An integral protein of the inner nuclear membrane localizes to the mitotic spindle in mammalian cells. Journal of Cell Science, 122, 2100-2107. http://dx.doi.org/10.1242/jcs.047373
[2] Ozcan, L., & Tabas, I. (2012). Role of Endoplasmic Reticulum Stress in Metabolic Disease and Other Disorders. Annual Review of Medicine, 63, 317-328. http://dx.doi.org/10.1146/annurev-med-043010-144749
[3] Vijayaraghavan, B., Jafferali, M. H., Fiqueroa, R. A., & Hallberg, E. (2016). Samp1, a RanGTP binding transmembrane protein in the inner nuclear membrane. Nucleus, 7(4), 415-423. http://dx.doi.org/10.1080/19491034.2016.1220465.